The clinical-stage company Verseon (Fremont, California) is unique among companies exploring the use of AI in drug discovery given its focus on what it calls the “uncharted” chemical space. While Big Pharma companies typically test compounds from their collections for specific diseases, these collections often have constraints and overlapping compounds. Adityo Prakash, CEO of Verseon, estimates that researchers have developed about seven million chemically distinct compounds globally in the past century. But compared to that figure, the number of mappable compounds based on organic synthesis principles is a staggering 1033.
Verseon is exploring uncharted chemical space

Prakash emphasizes that while most AI companies involved in drug discovery focus on the narrower subset contained in current libraries, Verseon has been systematically focused on exploring potentially more lucrative drug candidates plucked from the uncharted chemical space. “While AI is a buzzword now and people are trying to bring it into biotech, we’ve been at this, systematically building all the core fundamentals for a long, long time.”
Established in 2002, Verseon focuses on drug discovery programs for indications such as cardiometabolic diseases and cancer. The company employs a unique multiphysics-augmented AI platform, embedded in a comprehensive chemistry and biology workflow, to design new drug candidates for various diseases. Its pipeline includes VE-1902, a thrombin inhibitor for cardiovascular disorders in a phase 1 study.
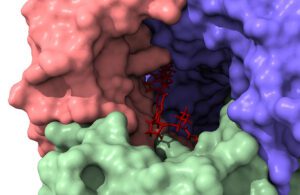
Instead of confining its focus to the current libraries, it focuses on the uncharted chemical space. “We determine chemical bond formations between specific atoms on the drug and the protein,” Prakash said. “We identify how all these atoms push and pull on each other, ranging from electrostatic interactions to potential hydrogen bond formations.”
In 2022, Verseon acquired Edammo, an AI company that developed a machine learning platform called Extreme AutoML, which outperforms Google AutoML in various life science tasks with a lower error rate. According to a research paper published by Springer, Extreme AutoML makes use of a neural architecture known as Extreme Learning Machines (ELMs) that tend to be less computationally expensive than deep learning. In tests, Edammo’s Extreme AutoML offered advantages in accuracy, Jaccard Indices, class variance and training times when benchmarked against Google’s AutoML using four popular classification datasets from the University of California at Irvine’s (UCI) repository. Prakash expects the Edammo acquisition to further enhance the company’s AI-based drug discovery platform leading to a more efficient and accurate drug discovery process.
Tackling the complexity problem in molecular modeling
Molecular modeling presents significant challenges owing to the inherent complexity of classical physics and quantum mechanics. “It is a fiendishly complicated problem to model interactions between a flexible target protein and a drug molecule in water, for instance,” Prakash said. “The problem lies in figuring out what will happen with a 10,000 atom protein interacting with a drug in the presence of tens of thousands of water molecules.”
Trying to brute force the core equations for quantum mechanics would overwhelm even the most powerful computers in the world. “New advancements in physics are needed to make the problem computationally tractable,” Prakash said.
Water molecules create a highly nonlinear problem that is difficult to model correctly. As a molecule binds, it vibrates, changes shape and experiences entropic effects. All of that complexity can vex researchers struggling to account for in their models. And understanding the behavior of discrete water molecules is also crucial for predicting how and where a drug molecule will bind.
“By developing in-house technologies to model these complex phenomena accurately, we are able to tackle challenges that others in academia, pharma or VC-backed companies might not be able to address,” Prakash said. “Our unconventional approach allows us to make progress in solving these complex problems, which lie at the heart of quantum mechanics and molecular modeling.”